- World
- Oct 09
3 scientists win Chemistry Nobel for work on proteins
• The Nobel Prize in Chemistry was awarded to three scientists for their breakthrough work predicting and even designing the structure of proteins, the building blocks of life.
• The prize was awarded to David Baker, who works at the University of Washington in Seattle, and to Demis Hassabis and John Jumper, who both work at Google DeepMind, a British-American artificial intelligence research laboratory based in London.
• David Baker has succeeded with the almost impossible feat of building entirely new kinds of proteins. Demis Hassabis and John Jumper have developed an AI model to solve a 50-year-old problem: predicting proteins’ complex structures.
• Half the prize was awarded to Baker for computational protein design, while the other half was shared by Hassabis and Jumper for protein structure prediction.
They cracked the code for proteins’ amazing structures
• The diversity of life testifies to proteins’ amazing capacity as chemical tools. They control and drive all the chemical reactions that together are the basis of life. Proteins also function as hormones, signal substances, antibodies and the building blocks of different tissues.
• Proteins are the molecules that enable life. Proteins are building blocks that form bones, skin, hair and tissue. To understand how life works, we first need to understand the shape of proteins.
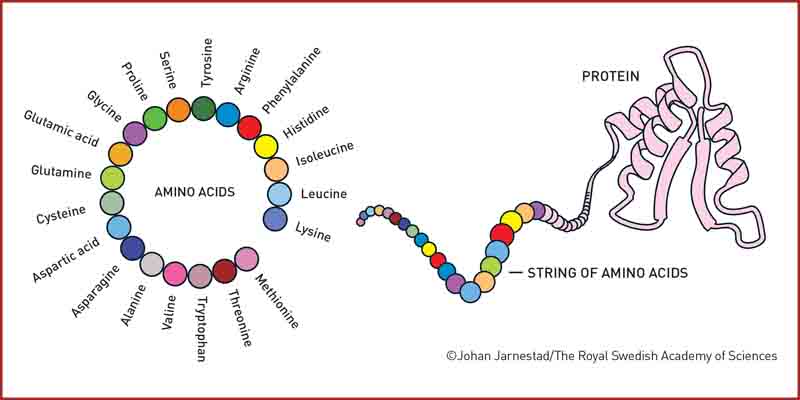
• Proteins generally consist of 20 different amino acids, which can be described as life’s building blocks. In 2003, David Baker succeeded in using these blocks to design a new protein that was unlike any other protein.
• Since then, his research group has produced one imaginative protein creation after another, including proteins that can be used as pharmaceuticals, vaccines, nanomaterials and tiny sensors.
• The second discovery concerns the prediction of protein structures. In proteins, amino acids are linked together in long strings that fold up to make a three-dimensional structure, which is decisive for the protein’s function. Since the 1970s, researchers had tried to predict protein structures from amino acid sequences, but this was notoriously difficult.
• In 2020, Demis Hassabis and John Jumper presented an AI model called AlphaFold2. With its help, they have been able to predict the structure of virtually all the 200 million proteins that researchers have identified.
• Google DeepMind has also made the code for AlphaFold2 publicly available, and anyone can access it. The AI model has become a gold mine for researchers. By October 2024, AlphaFold2 had been used by more than two million people from 190 countries.
• Previously, it often took years to obtain a protein structure, if at all. Now it can be done in a few minutes. The AI model is not perfect, but it estimates the correctness of the structure it has produced, so researchers know how reliable the prediction is.
A myriad of scientific applications
• Proteins’ amazing versatility as chemical tools is reflected in the vast diversity of life. That we can now so easily visualise the structure of these small molecular machines is mind boggling; it allows us to better understand how life functions, including why some diseases develop, how antibiotic resistance occurs or why some microbes can decompose plastic.
• The ability to create proteins that are loaded with new functions is just as astounding. This can lead to new nanomaterials, targeted pharmaceuticals, more rapid development of vaccines, minimal sensors and a greener chemical industry – to name just a few applications that are for the greatest benefit of humankind.
Manorama Yearbook app is now available on Google Play Store and iOS App Store